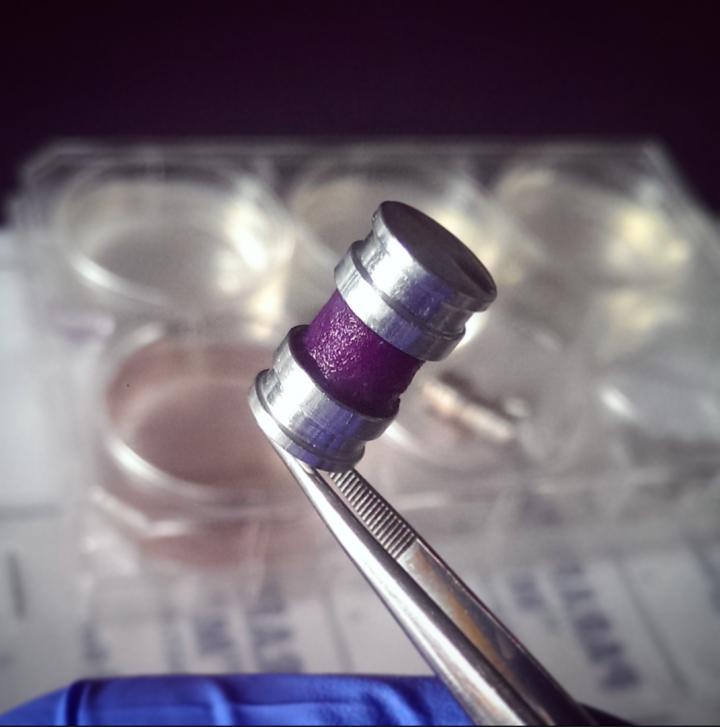
This rolled-up strip of engineered tissue allows scientists to mimic the way cells grow in a tumor, yet it can be unrolled in seconds for detailed analysis. (Credit: U of T Engineering/ Darren Rodenhizer)
A team of U of T engineers is unrolling the mysteries of cancer — literally. They have developed a way to grow cancer cells in the form of a rolled-up sheet that mimics the 3D environment of a tumor, yet can also be taken apart in seconds. The platform, described in a new Nature Materials paper, offers a way to speed up the development of new drugs and therapies and ask new questions about how cancer cells behave.
The drawbacks of studying cancer cells in a traditional petri dish are well known. While cells in a tumor grow in three dimensions, the dish is only two-dimensional. Moreover, cells in the center of a tumour have less access to oxygen and nutrients than those growing near the surface, close to the blood vessels. These subtle, location-dependent differences have a big impact on cell behavior, but have proven difficult to replicate in a dish.
“He wanted to do this analysis where you had to collect the cells in less than 10 seconds,” said Professor Alison McGuigan. (Credit: U of T Engineering/ Darren Rodenhizer)
In response, tissue engineers have tried to build more realistic 3D models by impregnating porous, sponge-like materials with cells and stacking them like building blocks. Chemical engineering professor Alison McGuigan is among them, but she was challenged to think differently about the problem by talking with fellow engineering professor Radhakrishnan Mahadevan.
“He wanted to do this analysis where you had to collect the cells in less than 10 seconds,” she says. “That was the engineering problem: how do you separate the cells in a very rapid way?” Building blocks wouldn’t do the trick; it would take too long to peel each layer off the structure and would be difficult to keep track of which block came from where.
McGuigan gathered materials from the lab (e.g. filter paper, petri dish, scissors, etc.) and started to think about how to assemble and disassemble them quickly. At one point, she absent-mindedly started wrapping a piece of paper around her finger. “I looked down at it, and realized that’s how you do it,” she says.
Over the next few years, McGuigan and her chemical engineering PhD student Darren Rodenhizer built their first prototype. They impregnated a short strip of a porous, paper-like support material with collagen — a gel-like material found in the body — and cancer cells. The whole thing was then bathed in a nutrient-rich culture solution for a day, allowing the cells to adjust to their new environment. Next, the strip was rolled around a metal core, forming an engineered tumor, which was then cultured for a few more days before performing analysis of tumor cell behavior.
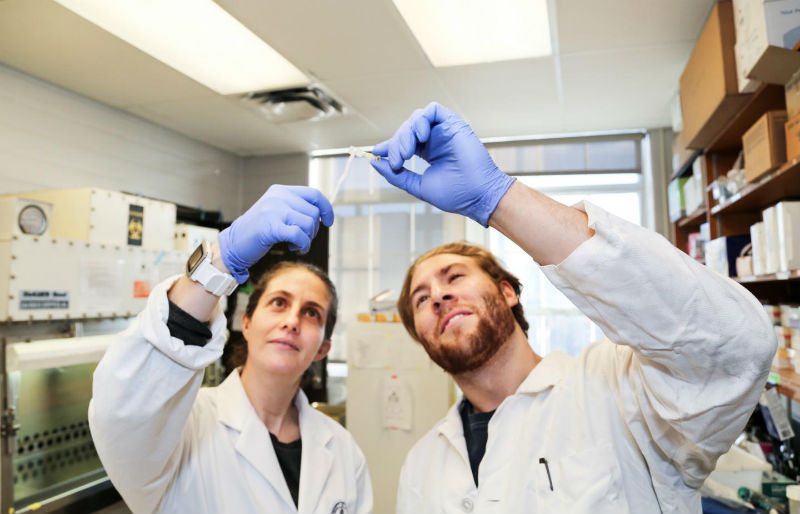

Professor Alison McGuigan and Ph.D. student Darren Rodenhizer unrolling the tumor. (Credit: U of T Engineering/ Ray Cheah)
Upon unrolling the device, the team found noticeable differences between the inner and outer layers. “As the oxygen level goes down, the number of dead cells in the layer increases, so the cells are responding to that oxygen gradient,” says Rodenhizer. Those cells that were still alive were shown to behave differently than the surface cells: for example, they more strongly expressed genes associated with low oxygen conditions. Crucially, the changes were gradual and continuous along the length of the strip. “If you had a stack, you could take it apart, but then you’d have all these separate, discontinuous pieces to keep track of,” says Rodenhizer “We have one layer.”
The single-layer design makes it easier for other lab researchers to adopt the process. “It’s simple enough that one could teach an undergrad to do it in a week,” says McGuigan. That makes it a boon to researchers looking to understand what makes cancerous cells in a tumor different from non-cancerous tissue. Exploiting these differences could accelerate the search for drugs that target cancer while leaving healthy cells alone.
Throughout the process, the team has collaborated closely with Christian Frezza at the University of Cambridge and Bradly G. Wouters at Princess Margaret Cancer Centre in Toronto to ensure that the tool enables the types of experimental tests that are most needed by cancer biologists to ask cutting edge questions and translate their findings into benefits to patients.
The technology also holds great promise for the field of personalized medicine. “The idea would be to take a patient’s own cells and create copies of their tumor,” says McGuigan. These copies could then be subjected to various treatments and analyzed by the simple unrolling process, providing information about what is likely to work best for that specific patient.
Now that it’s published, McGuigan hopes it will be widely adopted in the research community. “It’s very translatable and transferable to other labs,” says McGuigan. “We definitely want others to use it, because the larger the community, the more applications we will discover.”
