There are approximately 1,100 known antimicrobial peptides (AMP) with diverse sequences that can permeate microbial membranes. To help discover the “blueprint” for natural AMP sequences, researchers from the University of Illinois at Urbana-Champaign and the University of California, Los Angeles, have developed a new machine learning approach to discover and design alpha-helical membrane active peptides based on their physicochemical properties.
“In this work, we have trained a machine learning classifier–known as a support vector machine–to recognize membrane activity and experimentally calibrated the recognition metric by peptide synthesis and characterization,” explained Andrew Ferguson, an assistant professor of materials science and engineering at Illinois. “We use machine learning to not only discover new membrane active peptides, but to also identify membrane activity in known peptides with previously defined functions leading us to discover membrane activity in diverse and unexpected peptide families.
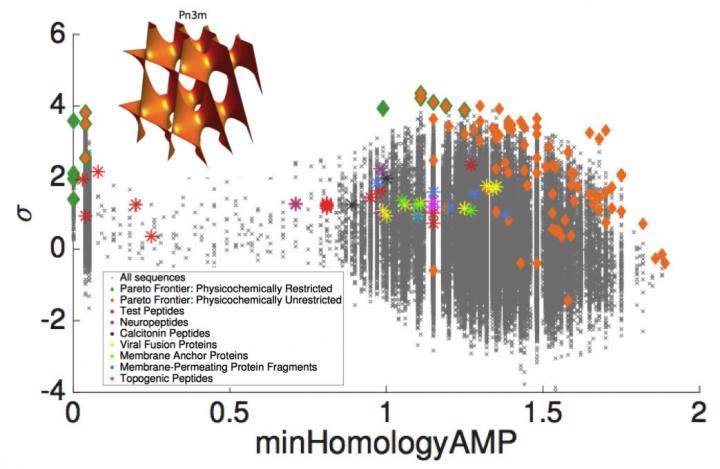
In this image the grey crosses are peptides scanned projected into a 2-D plane spanned by the sequence distance to a known antimicrobial peptide (x-axis) and prediction confidence of the classifier that the peptide is membrane active (y-axis). We detect membrane activity in diverse classes of peptides including neuropeptides, topogenic peptides, and viral fusion proteins. Membrane activity is mediated through the induction if negative Guassian curvature in the membrane (inset). (Credit: University of Illinois).
“Since getting cargo into a cell is important for many applications, we anticipate that this tool can have broad biomedical implications including in immunotherapy and in broad-spectrum membrane-active antimicrobial peptides to combat the rising incidence of drug resistance, design of cationic cell-penetrating peptides for nucleic acid transfection into cells, and in targeting and permeating anticancer therapeutics into tumors,” added Ferguson, who was the senior computational investigator for the project.
In this collaborative work, the Illinois researchers developed the computational innovations, with the experimental testing of the predictions accomplished at UCLA. The results, which highlight the difference between the efficacy of an antimicrobial and its recognizability as such, are surprising.
“AMPs do not share a common core structure, but tend to be short, cationic , and amphiphilic,” Ferguson said. “By training our machine learning classifier over a training set comprising peptides with known antimicrobial activity (hits) and decoy peptides with no activity (misses), the classifier learned the physical and chemical properties of a peptide that make for good membrane activity. We anticipated that the classifier would learn to discriminate the ‘antimicrobial-ness’ of a particular peptide sequence, but through experimental testing of its predictions we found that it actually learned a much more general and physical rule to discriminate peptides based on membrane activity. In effect, the classifier learned membrane activity as the underlying physical determinant of antimicrobial activity within the training set, and allows us to use our classifier to discover membrane active peptides in other diverse peptide classes.”
“Using the SVM as an efficient discovery tool for membrane activity, we performed a guided search of peptide sequence space to discover new membrane active peptides that would be difficult for nature to evolve by simple mutation from existing alpha-helical membrane active peptides.,” stated Ernest Y. Lee, first author of the paper, “Mapping membrane activity in undiscovered peptide sequence space using machine learning,” appearing in the Proceedings of the National Academy of Sciences.
“What emerges is a diverse taxonomy of sequences that are expected to be not only just as membrane-active as known antimicrobial peptides, but also have a broad range of putative primary functions beyond antimicrobial activity including neuropeptides, viral fusion proteins, topogenic peptides, and amyloids,” said Gerard Wong, a professor of bioengineering at UCLA and senior experimental investigator on the study. “Had their primary functions been undiscovered, these peptides could have been classified as AMPs. Not only is membrane activity not coextensive with antimicrobial activity, it is surprisingly common for many classes of natural peptides as one component of multiplexed functionality.”


