The facts about prostate cancer can be confusing. It’s the third most common cancer type among Americans — 161,000 men will be diagnosed this year, the National Cancer Institute estimates. Yet, according to the NCI, 98.6 percent will be alive in 5 years.
While it’s often not lethal, prostate cancer still kills lots of men — 21,000 this year — because it’s so common. Many of its victims have metastatic disease at the time of diagnosis — their cancer has already spread. Can we catch potentially lethal cancers earlier to save some of these men?
This week, a team at Cold Spring Harbor Laboratory (CSHL) publishes encouraging results of a pilot study testing a new way of pinpointing the minority who have aggressive disease at the time of diagnosis, a fact that usually determines whether they will have surgery or not. Current procedure calls for a predictive diagnostic biopsy, in which a needle is inserted into the organ at various locations, sometimes guided by MRI and ultrasound.
The new method — which is proposed as a way of augmenting pre-surgical biopsy results, not replacing them — uses advanced genomic tools to analyze what doctors call “biopsy cores.” In standard diagnostic biopsies, pathologists examine the tissue collected in these cores, usually a dozen, and assign an overall grade called a Gleason score, based on changes in glandular architecture. A score of 6 or lower is usually interpreted to mean a cancer is slow-growing or “indolent.” Yet some men with borderline Gleason scores undergo surgery and are found to have aggressive cancers. Other times, men with high Gleason scores are found to have indolent cancers upon surgery.
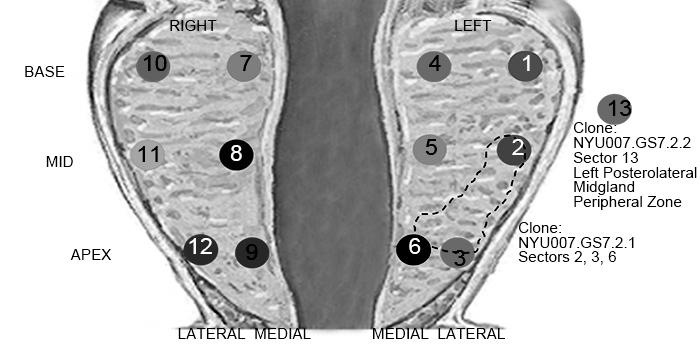
Rendering of the human prostate gland, roughly the size and shape of a walnut, bisected by the urethra. The locations of each of an individual’s 13 biopsy cores are indicated. Information obtained via single-cell sequencing from cells sampled from each core indicate the likely location of a cancerous tumor (dotted lines encompassing cores 2, 3 and 6). (Image credit: Krasnitz lab, CSHL)
The newly tested method, devised by CSHL Professor Michael Wigler and Associate Professor Alexander Krasnitz, draws the raw material for further analysis from the standard pre-surgical biopsy. The team sequences the genomes of several hundred single cells sampled from each patient’s biopsy cores. They search for certain patterns — for the presence of DNA disturbances called copy-number variations (CNVs). Using computational methods to compare CNV patterns, the team looks for cells whose CNV profiles harbor the same irregularities. This is a sign of clonality. Cancerous tumors are composed of clonal cells — genetically aberrant cells that derive from a single wayward ancestor.
The CSHL method assigns a number to each set of biopsy cores, based on how many of the cores contain clonal cells, how many clonal cells each contains, and how far such cells have dispersed within the prostate. In tests on 8 patients based on collaborations with NYU and Cornell University medical centers, the CSHL testing method yielded assessments of tumors that more closely matched the verdict of post-surgical pathological analysis (which reveal actual pathology) than the corresponding pre-surgical predictive biopsies, according to first author Joan Alexander, PhD.
“This is important because treatment decisions in such cases depend on the pre-surgical biopsy, not the surgical specimen,” Krasnitz comments. “We think single-cell analysis could potentially augment traditional biopsy-core histopathology, significantly improving risk assessment and informing treatment decisions, especially in borderline cases.”