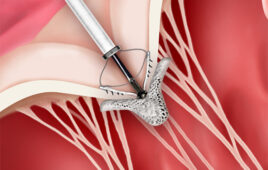
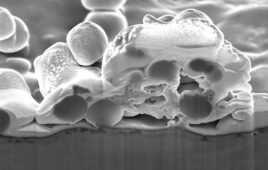
Solar thermal PCR system: (a), the benchtop component holds the lens, the PCR chip and a rotational stage. (b), Thermocouples on the chip are connected by a microcontroller to a smartphone. (c), a screenshot shows the app recording the temperatures over time. (d), the top of the PCR chip contains the mask and the thermocouples. (e), the bottom includes the absorber layer and the microfluidic channel, with marked regions for annealing, extension and denaturation (inset). (f), amplification over a range of concentrations. (g), amplification over a range of cycling speeds. The typical extension rate of Taq polymerase is 60–100 nucleotides/s at 72°C. Thus, 3 s should be sufficient for full extension of a 164 bp product. The design of our channel suggests that a minimum reaction time of 10 s/cycle is required. The band intensity began decreasing for reaction times faster than 20 s/cycle, and no band was observed for reactions faster than 10 s/cycle. Intensity values were normalized by a reference sample that was run in a conventional thermal cycler for 2 h.

Thermal characterization and demonstration of solar thermal PCR: (a), schematic showing the ability to change the solar intensity by changing the lens-to-chip distance. (b), a simulation showing the plateaued temperature profile at the plane of the microchannel. (c), measurements in (i) April and (ii) May demonstrate the ability to achieve similar on-chip temperatures using the same mask. (iii) Simulations provide thermal profiles from 0°C to 30°C ambient temperature. (d), Thermal measurements from 7 AM to 7 PM show relatively consistent values. (e), averaged temperature data shows day-long trend. f, PCR tests show that hourly changes in ambient temperature still allowed for product amplification. For Fig. 2c, d, and e, the red, green, and blue color curves respectively refer to the denaturation, extension, and annealing temperatures measured in the app.

Characterization of simulated cloud coverage and flow rate: (a), temperature variation for denaturation, extension and annealing due to blocking of the light. Simulated clouding times ranged from 0 s (darkest curve) to 4 min (lightest curve) for a total test time of 27 min. The percentages represent the percent of time that each test spent in non-ideal PCR conditions, which for these tests was defined as when the denaturation temperature dropped below the dashed line. (b), gel electrophoresis and (c), corresponding band measurements show diminishing intensities as a function of the duration of simulated cloud coverage.

Integration of HotSHOT cell lysis, solar thermal PCR and smartphone-based detection applied to mouse tail biopsies: (a), fluorescence detection setup includes a blue filter between the LED and the chip and a green filter between the chip and the camera. (b), PDMS chip (inset) containing 4 tests: solar thermal PCR performed using KSHV+ samples (1, 2) and a KSHV- sample (3) and traditional PCR using negative control (NC). (c), a screenshot of the app that analyzes the fluorescence signals, showing high intensities for samples 1 and 2 and low intensities for sample 3 and NC.

Li Jiang, a doctoral candidate in engineering, explains the solar thermal PCR test for a smartphone. The system can detect Kaposi’s sarcoma – a deadly skin cancer related to HIV infection – in about a half-hour. (Credit: Jason Koski/University Photography)

A smartphone application, working in conjunction with the PCR device, provides results of the diagnostic tests. (Credit: Jason Koski/University Photography)