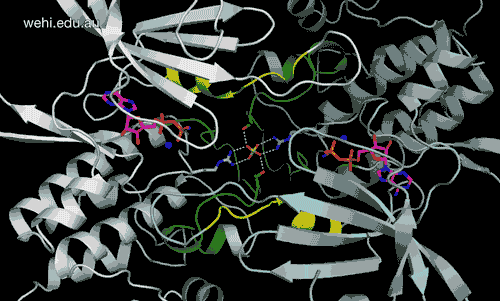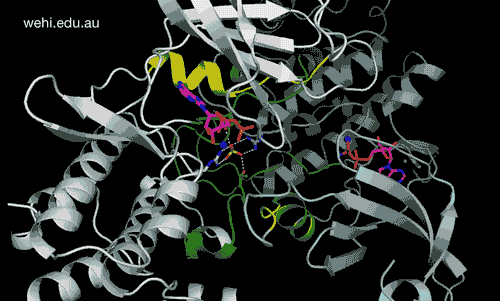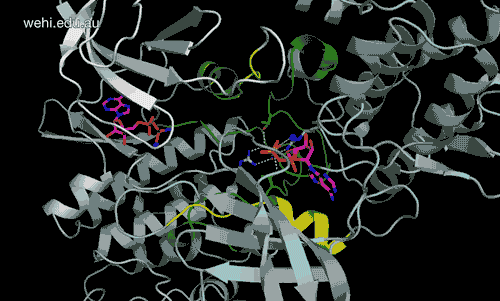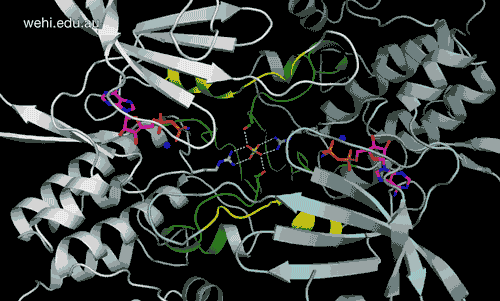
Walter and Eliza Hall Institute researchers have generated the first 3-D map of the cancer-associated protein DCLK1, revealing how it functions. Credit: Walter and Eliza Hall Institute, Australia
Walter and Eliza Hall Institute researchers have revealed for the first time the three-dimensional molecular ‘map’ of a protein that has been pinpointed as a driver of many types of cancers.
The unprecedented view of the protein doublecortin kinase like domain 1 (DCLK1) could provide clues to how it contributes to cancer formation and progression.
DCLK1 is a protein that assembles scaffolds within cells called microtubules. These rope-like structures give cells shape, enable movement and cell division, and are crucial in enabling the growth and spread of cancer cells. More than one in 10 stomach cancers have defective forms of DCLK1, which have also been found in kidney, rectal and pancreatic cancers.
Walter and Eliza Hall Institute scientists Dr Onisha Patel and Dr Isabelle Lucet used the Australian Synchrotron to reveal the three-dimensional structure of a part of DCLK1 known as the ‘kinase domain’.
Working with Professor Matthias Ernst and Dr Michael Buchert at the Olivia Newton-John Cancer Research Institute, the team was able to pinpoint the parts of DCLK1 that can go wrong in cancer cells, providing new information about how the protein functions. The research is published today in the journal Structure.
Dr Lucet said in healthy cells, DCLK1 can control its own function, ensuring it only assembles microtubules at the right time. “We can think of the kinase domain of DCLK1 being an inbuilt controlling unit for the protein,” she said. “The kinase domain sits separately from the part of the protein that assembles microtubules, and can switch microtubule assembly on or off as needed.
“The complexity of having these two different components in one protein has, until now, hindered our understanding of how DCLK1 functions normally and what goes wrong in cancer. We decided to focus on just the regulatory kinase domain of DCLK1, because we knew that this is the part of DCLK1 that is often altered in cancers,” Dr Lucet said.
The team created a detailed map of the DCLK1 kinase domain using the Australian synchrotron, providing new information about how the part of the protein functioned, Dr Patel said.
“By looking at the structure of the kinase domain at the atomic level, we can now understand the effect of cancer-associated mutations located in this region,” she said. “These mutations can lead to a non-stop microtubule building by DCLK1, and consequently chaotic cell division that is a hallmark of cancer.
“The protein structure we have generated allows us to pinpoint a crucial three dimensional pocket within DCLK1. We hope this will provide a first step towards designing a drug that will precisely target DCLK1, and prevent it from driving cancer growth. Similar protein structures have been used to design anti-cancer agents that are now in clinical use, so we’re hopeful that our research may have similar applications into the future,” Dr Patel said.
Explore further: Researchers unlock secrets of troublesome Tribble protein
More information: DOI: 10.1016/j.str.2016.07.008
