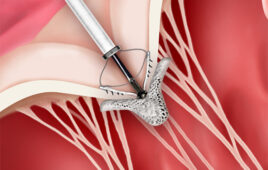
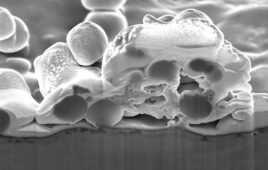
Cryo-electron tomography permits researchers to study in detail the microscopic structures inside of cells. Researchers who typically required a week of effort to dissect the 3-D structure of a single cell will now be able to do it in about an hour thanks to a new automated method developed by a team of scientists at Baylor College of Medicine and the National University of Singapore. The new method will allow scientists to study a large number and a variety of cell types in significantly less time, leading to a more detailed understanding of cellular processes and disease. The report appears in the journal Nature Methods.

This is Dr. Steven Ludtke. (Credit: Baylor College of Medicine)
“Cryo-electron tomography is a powerful technology for visualizing the architecture and structures inside of cells at about 100 times better resolution than that of the best light microscope,” said corresponding author Dr. Steven Ludtke, co-director of the National Center for Macromolecular Imaging and the Center for Computational and Integrative Biomedical Research and professor of biochemistry at Baylor College of Medicine.
Cryo-electron tomography provides information about cellular processes that cannot be obtained by any other current method. Scientists can study changes inside normal or diseased cells as well as inside microorganisms. However, “the need for about one man-week of effort to manually annotate all of the structures and details within each cell has limited the widespread use of this technique,” Ludtke said.
Looking to increase the efficiency of the time-consuming process of annotation, Ludtke and his colleagues have developed an automated method that requires less human participation. They tested their method using different cell types, including human platelets and microorganisms such as
African trypanosomes and cyanobacteria. When they compared the new method with the traditional, they found that the new methodology correctly identified structures inside cells with a high level of accuracy.
“The new method reduces human effort from about one week per cell to about one hour per cell,” Ludtke said. Researchers interested in applying this method will find it available at no cost at http://EMAN2.org, together with an interface for training.