Research team led by Nagoya University develop a label-free method for detecting DNA amplification in real time based on refractive index changes in diffracted light.
Polymerase chain reaction (PCR) is a simple and ubiquitous method in molecular biology for amplifying DNA segments into millions of copies. This is important not only for basic research, but also in diagnostics, forensics, and medical applications. Quantitative real-time PCR is a modified version that incorporates fluorescence labeling to cumulatively measure DNA amplification, rather than monitoring it at the end of the process, as in conventional PCR. Real-time PCR therefore enables sensitive quantification of the amount of the initial DNA template. However, current techniques may introduce bias through sequence errors, pipetting inaccuracies, or unequal binding of fluorescent probes (hybridization).
A research team centered on Nagoya University has now developed a novel method of measuring real-time DNA amplification that is label-free, thus avoiding the bias issues associated with other procedures. The research and its outcomes were reported in Scientific Reports.
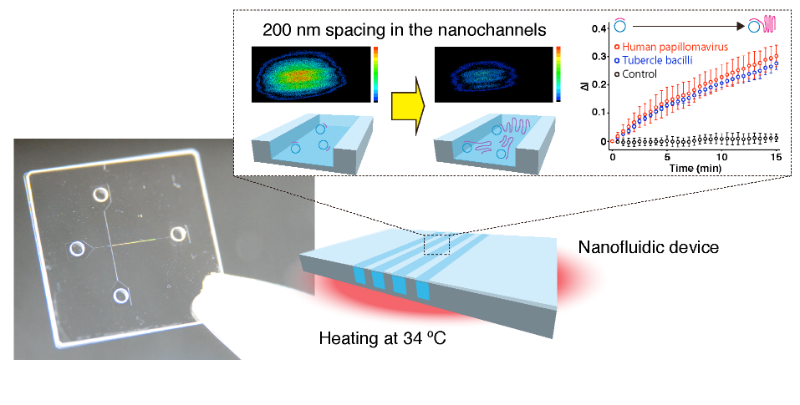
A photo and a schematic illustration for a nanofluidic diffraction grating. Label-free signals based on a diffraction intensity change were attributed to amplification of DNA molecules, such as human papillomavirus and tubercle bacilli. (Credit: Copyright Takao Yasui)
Existing label-free detection systems rely on surface immobilization of target molecules, which is expensive, laborious, and ineffective over time. This new method also introduces an element of hybridization bias because of complementary probe binding. The new technique instead detects changes in the intensity of diffracted light from a laser beam passing through miniscule 200 nm (0.0002 mm)-wide nanochannels filled with analytical sample liquids. The 532-nm laser beam is focused by a lens and then diffracted by passing through the nanochannel and detected by a photodiode. Silica substrates were used to make the nanochannels, and the larger the difference between refractive indices of sample liquids and silica, the smaller is the change in diffracted light intensity, and vice versa.
“We used this technique to provide the first label-free detection of human papillomavirus and the bacteria responsible for tuberculosis,” first author Takao Yasui says. “The method is highly sensitive, and allows quantification of a wide range of initial DNA concentrations, from 1 fM to 1 pM (a 1,000-fold range), so is superior to existing fluorescence-based detection systems.”
“Our system also measures DNA amplification at the relatively low temperature of 34°C and without the need for thermal cycles,” coauthor Noritada Kaji says. “Because it has the potential to be constructed as a single chip and can detect sample volumes as small as 1 µl, which is 100–1,000 times less than conventional detectors are capable of, it is particularly suited to development as a miniaturized form of diagnostics and microbe detection.”
(Source: AlphaGalileo)