Newest method for determining a protein’s shape based on XFEL technology significantly broadens number and type of proteins that researchers can study
 In biology, a protein’s shape is key to understanding how it causes disease or toxicity. Researchers who use X-rays to take snapshots of proteins need a billion copies of the same protein stacked and packed into a neat crystal. Now, scientists using exceptionally bright and fast X-rays can take a picture that rivals conventional methods with a sheet of proteins just one protein molecule thick.
In biology, a protein’s shape is key to understanding how it causes disease or toxicity. Researchers who use X-rays to take snapshots of proteins need a billion copies of the same protein stacked and packed into a neat crystal. Now, scientists using exceptionally bright and fast X-rays can take a picture that rivals conventional methods with a sheet of proteins just one protein molecule thick.
Using a type of laser known as XFEL, the technique opens the door to learning the structural details of almost 25 percent of known proteins, many of which have been overlooked due to their inability to stack properly. The team of researchers led by the Department of Energy’s Pacific Northwest and Lawrence Livermore National Laboratories report their results with this unique form of X-ray diffraction in the March issue of the International Union of Crystallography Journal.
“In this paper, we’re proving it’s possible to use an XFEL to study individual monolayers of protein,” said PNNL microscopist James Evans. “Just being able to see any diffraction is brand new.”
Evans co-led the team of two dozen scientists with LLNL physicist Matthias Frank. The bright, fast X-rays were produced at the Linac Coherent Light Source at SLAC National Accelerator Laboratory in Menlo Park, Calif., the newest of DOE’s major X-ray light source facilities at the national laboratories. LCLS, currently the world’s most powerful X-ray laser, is an X-ray free-electron laser. It produces beams millions of times brighter than earlier X-ray light sources.
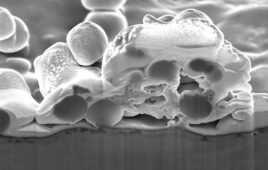
Coming in at around 8 angstrom resolution (which can make out items a thousand times smaller than the width of a hair), the proteins appear slightly blurry but match the expected view based on previous research. Evans said this level of clarity would allow researchers, in some cases, to see how proteins change their shape as they interact with other proteins or molecules in their environment.
To get a clearer view of protein monolayers using XFEL, the team will need to improve the resolution to 1 to 3 angstroms, as well as take images of the proteins at different angles, efforts that are currently underway.
Not Your Family’s Crystal
Researchers have been using X-ray crystallography for more than 60 years to determine the shape and form of proteins that form the widgets and gears of a living organism’s cells. The conventional method requires, however, that proteins stack into a large crystal, similar to how oranges stack in a crate. The structure of more than 80,000 proteins have been determined this way, leading to breakthroughs in understanding of diseases, pathogens, and how organisms grow and develop.
But many proteins found in nature do not stack easily. Some jut from the fatty membranes that cover cells, detecting and interacting with other cells and objects, such as viruses or bacteria, in the surrounding area. These proteins are not used to having others of their kind stack on top. These so-called membrane proteins make up about 25 percent of all proteins but only 2 percent of proteins that researchers have determined structures for.
Wafer Thin
Researchers in the last decade have been pursuing the idea that one sheet of proteins could be visualized if the X-rays were bright enough but flashed on and off quickly enough to limit the damage wrought by the powerful X-rays. Two years ago, scientists demonstrated they could use XFEL technology on crystals of proteins about 15 to 20 sheets thick.
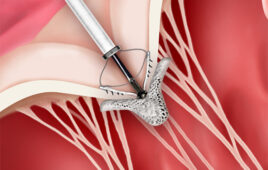
Evans, Frank and their team wanted to push this further. The team worked on a way to create one-sheet-thick crystals of two different proteins — a protein called streptavidin and a membrane protein called bacteriodopsin. The structures of both proteins are well-known to scientists, which gave the team something to compare their results to.
The team shined the super-bright X-rays for a brief moment — about 30 femtoseconds, a few million billionths of a second — on the protein crystals. They created so much data in the process that it took them more than a year to analyze all of it.
The resulting images look like the known structures, validating this method. Next, the researchers will try to capture proteins changing shape as they engage in a chemical reaction. For this, even shorter flashes of X-rays might be needed to see the action clearly.
If successful, shorter flashes of XFEL might mean longer lines at SLAC.
Evans is based at EMSL, the Environmental Molecular Sciences Laboratory at PNNL. This work was supported by the Department of Energy Office of Science, National Institutes of Health, National Science Foundation, Lawrence Livermore National Laboratory, and Pacific Northwest National Laboratory.
For more information, visit Pacific Northwest National Laboratory.